If you haven't been reading my last entries, I have been working on a program that tries to design a circuits using genetic algorithm that fulfill the requirements given by the programmer. Read this entry for an overview what the program does and how it works. In this post I will tell about my findings about which algorithms and parameters work and which don't.
Population size
Before I started writing this program I read some literature about similar experiments. For example I read some John R. Koza's books about using genetic algorithms on designing electric circuits and he used some huge population sizes. In the book Genetic Programming IV: Routine Human-Competitive Machine Intelligence population sizes that he uses are usually tens of thousands. Some of the runs use population size of 100000 which is absolutely huge. This helps to explore the whole search space and guarantees a good solution. But I have found that population sizes of this big are unnecessary and they slow down the simulation too much. I have experimented and found that population sizes of 100-2000 are often good enough and they convergence a lot of quicker to very similar solutions as the population sizes of 10000 to 20000. This is often a difference between a few hour simulation and a few days of simulation.
Crossover
I have found that the way I currently do crossovers, by cutting and pasting parts of the two circuits netlist to one new netlist often produces bad results and I have been thinking that the whole existence of crossovers might be just waste and it would be a better use of resources to just disable them. Currently on my latest files I have set the crossover probability to low value of 1% to 5%, compared to mutation probability of 70%.
The problem with crossovers is that by applying it to two similar circuits we get one circuit that has many similar parallel devices. If this circuit with parallel devices performs better than the previous circuit parallel devices will soon spread within the population and they decrease the number of devices available to the rest of the circuit. Getting rid of the unnecessary parallel device, for example a two resistor requires two lucky mutations: one of the resistors needs to be removed and others value needs to be halved. Probability of this happening is small and instead it would have been better if in the first place there would have been a mutation in the value of the resistor that would have decreased it's value. This way it wouldn't have introduced an unnecessary device and it could have been used in a better way.
Circuit encoding
I decided to encode the circuits as a simple SPICE netlists. The problem with them is that crossovers often introduce duplicate parallel devices and seem harm more than help. Also they lack clear input and output. It is equally likely that a new device will connect straight from input to output or it will connect somewhere else. This usually leads to circuits that are not very deep, meaning that input and output nodes are very close together. This makes hard to evolve circuits that need to be deep to work correctly, for example higher order filters are necessarily deep.
Koza uses binary trees and set of functions to modify it. I have not tried them myself, but they don't suffer from same problems as netlists. Crossover is more efficient because subtrees can be exchanged. It's also easier to evolve deeper circuits when whole subtrees can be moved around.
An alternatives to these two encodings would be a graph based encoding. Graphs would work by having a starting node and a list of instructions that insert devices, change the current node and instruction that does both. This way duplicating a piece of genome makes a new series connected stage that is connected to previous stage instead of duplicating devices of the previous stage. I have not tried this approach yet. It's interesting, but it has some problems. Coding it is harder than simple netlists, it is more computationally intensive because the circuit needs to be converted to netlist for simulation and handling devices with more than two connections(e.g. Transistors) will be harder. Hopefully it will be worth it and evolving deeper circuits will become easier.
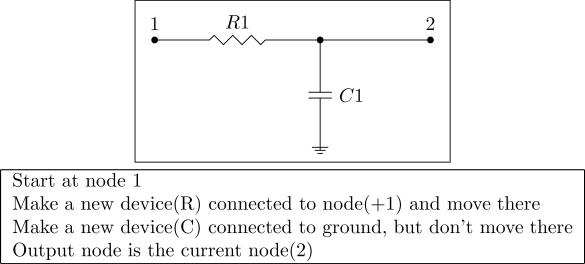
Lowpass filter encoded with graph encoding.
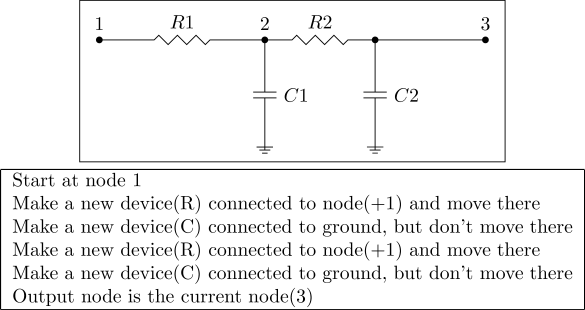
Same lowpass filter, but with a crossover that duplicated the instructions. Instead of parallel devices, the circuit now has a second stage.
Scoring
One of the most important things I have found that improve the quality of evolved circuits is that instead of keeping the constraint functions scores constant I ramp up the scoring weights over many generations and increase the strictness of constraints. For example when evolving an inverter I had a problem where my constraints for the current use were so strict that the program decided that it's better to not make anything to keep current use low instead of trying to make an inverter and use current. Ramping constraint scores solves this problem by allowing the program first make circuits that use lots of current, but work like they should and after some working circuits are found concentrate on fulfilling the constraints. In the inverter example this leads to very current efficient inverters that work like they should.
Summary
Since announcing this project I have made many improvements. Current and used power can be measured and controlled. Better scoring and improvements in scoring algorithm. New mutations that can change the output node and other mutation that changes a value of a device only a little amount, probably other mutations that I have forgotten already. Device can have an arbitrary cost(price, amount of area used...) that is added to the circuits score. The biggest improvement might be the gradual ramping of constraint scores that generate higher quality solutions. I have also replaced the unmaintainable way of writing the circuits straight to the programs source code with separate configuration files that are easier to write and maintain.
I have also added more options. For example it is now possible to not use the default scoring functions, but instead to write your own function that takes the simulation output and returns the score of the circuit.
I have also since replaced Python with PyPy. It uses much less memory, usually two to ten times less. With extremely big populations memory saving are even bigger. There might be something weird in the Python reference counting that keeps useless simulation results in memory instead of freeing them. I haven't investigated it much, because PyPy works so much better. Only problem with PyPy is that matplotlib isn't supported. Currently I run matplotlib as a subprocess, but PyPy team is working on supporting it and I hope that soon I can do the plotting also with PyPy.